Note
This notebook is located in the ./examples directory of the gwtransport repository.
Aquifer Characterization Using Temperature Response#
Learning Objectives#
Understand how temperature can be used as a natural tracer for aquifer characterization
Learn inverse modeling techniques for estimating aquifer properties
Apply gamma distribution models to represent aquifer heterogeneity
Interpret thermal breakthrough curves in hydrogeological contexts
Overview#
This notebook demonstrates inverse modeling to estimate aquifer pore volume distribution from temperature breakthrough curves. Temperature acts as a conservative tracer with known thermal retardation, allowing characterization of flow paths and residence times. Transport is modeled with the diffusion_fast module, which accounts for advection, microdispersion, and molecular diffusion.
Applications#
Groundwater vulnerability assessment
Residence time distribution analysis
Contaminant transport forecasting
Aquifer heterogeneity characterization
Key Assumptions#
Stationary pore volume distribution (steady streamlines)
Thermal retardation factor = 2.0 (typical for saturated media)
Transport includes advection, microdispersion, and molecular diffusion via
diffusion_fast
Background Reading#
Pore Volume Distribution — Central concept for aquifer heterogeneity modeling
Retardation Factor — How sorption/thermal exchange slows transport
Gamma Distribution — Two-parameter model for pore volumes
Dispersion Scales — Macro vs microdispersion
Theoretical Background#
Thermal Transport in Groundwater#
The residence time for thermal transport relates to the aquifer pore volume distribution through:
where \(V_{pore}\) is the pore volume [m³], \(R_f\) the thermal retardation factor [-], and \(Q\) the flow rate [m³/day]. See residence time.
Gamma Distribution Model#
Aquifer heterogeneity is represented using a gamma distribution for pore volumes, characterized by shape (\(\alpha\)) and scale (\(\beta\)) parameters, or equivalently by mean and standard deviation.
See concepts and assumptions for further background.
[1]:
import matplotlib.pyplot as plt
import numpy as np
from scipy.optimize import curve_fit
from scipy.stats import gamma as gamma_dist
from gwtransport import diffusion_fast
from gwtransport import gamma as gamma_utils
from gwtransport.examples import generate_temperature_example_data
from gwtransport.residence_time import fraction_explained, residence_time
from gwtransport.utils import step_plot_coords
# Set random seed for reproducibility
np.random.seed(42)
plt.style.use("seaborn-v0_8-whitegrid")
print("Libraries imported successfully")
Libraries imported successfully
1. Synthetic Data Generation#
We generate realistic temperature and flow time series to demonstrate the inverse modeling approach. The synthetic data includes seasonal patterns and realistic variability.
[2]:
# Generate 6 years of daily data with seasonal patterns
df, tedges = generate_temperature_example_data(
date_start="2020-01-01",
date_end="2025-05-31",
flow_mean=120.0, # Base flow rate [m³/day]
flow_amplitude=40.0, # Seasonal flow variation [m³/day]
flow_noise=5.0, # Random daily fluctuations [m³/day]
cin_method="soil_temperature", # Use real soil temperature data
aquifer_pore_volume_gamma_mean=8000.0, # True mean pore volume [m³]
aquifer_pore_volume_gamma_std=400.0, # True standard deviation [m³]
measurement_noise=0.1, # Measurement noise for temperatures. Set to zero for perfect fit.[°C]
)
print("Dataset Summary:")
print(f"Period: {df.index[0].date()} to {df.index[-1].date()}")
print(f"Mean flow: {df['flow'].mean():.1f} m³/day")
print(f"Mean infiltration temperature: {df['cin'].mean():.1f} °C")
print(f"Mean extraction temperature: {df['cout'].mean():.1f} °C")
print(f"True mean pore volume: {df.attrs['aquifer_pore_volume_gamma_mean']:.1f} m³")
print(f"True std deviation: {df.attrs['aquifer_pore_volume_gamma_std']:.1f} m³")
Dataset Summary:
Period: 2020-01-01 to 2025-05-31
Mean flow: 110.6 m³/day
Mean infiltration temperature: 11.8 °C
Mean extraction temperature: 12.9 °C
True mean pore volume: 8000.0 m³
True std deviation: 400.0 m³
2. Parameter Estimation via Optimization#
We implement inverse modeling to estimate gamma distribution parameters using nonlinear least squares optimization. A spin-up period is excluded to allow thermal breakthrough to stabilize. We use :func:~gwtransport.residence_time.fraction_explained to compute the fraction of pore volumes with valid residence times, and exclude the period where the model output is not yet fully informed by the input signal.
[3]:
# Compute fraction_explained to determine the spin-up period.
# We use the initial parameter guesses (p0) to estimate when the model output
# is fully informed by the input signal. After fitting, we verify that the
# spin-up period was sufficient.
p0_mean, p0_std = 7500.0, 450.0
p0_bins = gamma_utils.bins(mean=p0_mean, std=p0_std, n_bins=25)
rt = residence_time(
flow=df.flow,
flow_tedges=tedges,
aquifer_pore_volumes=p0_bins["expected_values"],
retardation_factor=2.0, # Thermal retardation
direction="extraction_to_infiltration",
)
frac = fraction_explained(rt=rt)
# Find the first date where the model output is fully explained
fully_explained_mask = frac >= 0.99
first_fully_explained = df.index[fully_explained_mask][0]
# Define training dataset excluding the spin-up period
train_data = df.loc[fully_explained_mask].cout
train_data = train_data.dropna()
train_length = len(train_data)
print(f"Spin-up period ends: {first_fully_explained.date()}")
print(f"Training dataset: {train_length} days")
print(f"Training period: {train_data.index[0].date()} to {train_data.index[-1].date()}")
Spin-up period ends: 2020-08-03
Training dataset: 1763 days
Training period: 2020-08-03 to 2025-05-31
[4]:
def objective(_xdata, mean, std):
"""Infiltration to extraction model for temperature breakthrough with gamma-distributed pore volumes."""
print(f"Optimizing: mean={mean:.1f} m³, std={std:.1f} m³")
cout = diffusion_fast.gamma_infiltration_to_extraction(
cin=df.cin,
flow=df.flow,
tedges=tedges,
cout_tedges=tedges,
mean=mean, # Mean pore volume [m³]
std=std, # Standard deviation [m³]
n_bins=25, # Discretization resolution
retardation_factor=df.attrs["retardation_factor"], # Thermal retardation factor
mean_longitudinal_dispersivity=df.attrs["longitudinal_dispersivity"], # Longitudinal dispersivity [m]
mean_molecular_diffusivity=df.attrs["molecular_diffusivity"], # Molecular diffusion coefficient [m²/day]
mean_streamline_length=df.attrs["streamline_length"], # Streamline length [m]
)
# Return training period data (excluding spin-up)
return cout[fully_explained_mask]
[5]:
# Nonlinear least squares optimization
print("Starting parameter optimization...")
(mean, std), pcov = curve_fit(
objective,
df.index,
train_data.values,
p0=(7500.0, 450.0), # Initial parameter estimates [m³]
bounds=([5000, 200], [10000, 600]), # Physical constraints [m³]
method="trf", # Trust Region Reflective algorithm
max_nfev=250, # Limit function evaluations
)
print("\nOptimization completed!")
Starting parameter optimization...
Optimizing: mean=7500.0 m³, std=450.0 m³
/Users/bdestombe/Projects/gwtransport/gwtransport/src/gwtransport/diffusion_fast.py:596: UserWarning: Using multiple aquifer_pore_volumes with non-zero mean_longitudinal_dispersivity. Both represent spreading from velocity heterogeneity at different scales.
This is appropriate when:
- APVD comes from streamline analysis (explicit geometry)
- You want to add unresolved microdispersion and molecular diffusion
This may double-count spreading when:
- APVD was calibrated from measurements (microdispersion already included)
For gamma-parameterized APVD, consider using the 'equivalent APVD std' approach
from 05_Diffusion_Dispersion.ipynb to combine both effects in the faster advection
module. Note: this variance combination is only valid for continuous (gamma)
distributions, not for discrete streamline volumes.
Suppress this warning with suppress_dispersion_warning=True.
return infiltration_to_extraction(
Optimizing: mean=7500.0 m³, std=450.0 m³
Optimizing: mean=7500.0 m³, std=450.0 m³
Optimizing: mean=7893.0 m³, std=477.1 m³
Optimizing: mean=7893.0 m³, std=477.1 m³
Optimizing: mean=7893.0 m³, std=477.1 m³
Optimizing: mean=7989.0 m³, std=449.0 m³
Optimizing: mean=7989.0 m³, std=449.0 m³
Optimizing: mean=7989.0 m³, std=449.0 m³
Optimizing: mean=8001.6 m³, std=427.1 m³
Optimizing: mean=8001.6 m³, std=427.1 m³
Optimizing: mean=8001.6 m³, std=427.1 m³
Optimizing: mean=8002.3 m³, std=419.2 m³
Optimizing: mean=8002.3 m³, std=419.2 m³
Optimizing: mean=8002.3 m³, std=419.2 m³
Optimizing: mean=8002.8 m³, std=413.9 m³
Optimizing: mean=8002.8 m³, std=413.9 m³
Optimizing: mean=8002.8 m³, std=413.9 m³
Optimizing: mean=8003.1 m³, std=408.7 m³
Optimizing: mean=8003.1 m³, std=408.7 m³
Optimizing: mean=8003.1 m³, std=408.7 m³
Optimizing: mean=8002.6 m³, std=405.8 m³
Optimizing: mean=8002.6 m³, std=405.8 m³
Optimizing: mean=8002.6 m³, std=405.8 m³
Optimizing: mean=8002.7 m³, std=401.9 m³
Optimizing: mean=8002.7 m³, std=401.9 m³
Optimizing: mean=8002.7 m³, std=401.9 m³
Optimizing: mean=8002.9 m³, std=396.1 m³
Optimizing: mean=8002.9 m³, std=396.1 m³
Optimizing: mean=8002.9 m³, std=396.1 m³
Optimizing: mean=8002.5 m³, std=392.5 m³
Optimizing: mean=8002.5 m³, std=392.5 m³
Optimizing: mean=8002.5 m³, std=392.5 m³
Optimizing: mean=8002.6 m³, std=388.8 m³
Optimizing: mean=8002.6 m³, std=388.8 m³
Optimizing: mean=8002.6 m³, std=388.8 m³
Optimizing: mean=8002.6 m³, std=387.0 m³
Optimizing: mean=8002.6 m³, std=387.0 m³
Optimizing: mean=8002.6 m³, std=387.0 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.6 m³, std=387.2 m³
Optimizing: mean=8002.6 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimizing: mean=8002.5 m³, std=387.3 m³
Optimization completed!
[6]:
# Generate model predictions using optimized parameters
df["cout_modeled"] = diffusion_fast.gamma_infiltration_to_extraction(
cin=df.cin,
flow=df.flow,
tedges=tedges,
cout_tedges=tedges,
mean=mean, # Fitted mean pore volume
std=std, # Fitted standard deviation
n_bins=250, # High computational resolution
retardation_factor=df.attrs["retardation_factor"], # Thermal retardation
mean_longitudinal_dispersivity=df.attrs["longitudinal_dispersivity"], # Longitudinal dispersivity [m]
mean_molecular_diffusivity=df.attrs["molecular_diffusivity"], # Molecular diffusion coefficient [m²/day]
mean_streamline_length=df.attrs["streamline_length"], # Streamline length [m]
)
# Report optimization results with uncertainty estimates
print("Parameter Estimation Results:")
print(f"Mean pore volume: {mean:.1f} ± {pcov[0, 0] ** 0.5:.1f} m³")
print(f"Standard deviation: {std:.1f} ± {pcov[1, 1] ** 0.5:.1f} m³")
print(f"Coefficient of variation: {std / mean:.2f}")
# Compare with true values
true_mean = df.attrs["aquifer_pore_volume_gamma_mean"]
true_std = df.attrs["aquifer_pore_volume_gamma_std"]
print(f"\nTrue values: {true_mean:.1f} m³ (mean), {true_std:.1f} m³ (std)")
print(
f"Relative error: {abs(mean - true_mean) / true_mean * 100:.1f}% (mean), {abs(std - true_std) / true_std * 100:.1f}% (std)"
)
Parameter Estimation Results:
Mean pore volume: 8002.5 ± 1.6 m³
Standard deviation: 387.3 ± 4.2 m³
Coefficient of variation: 0.05
True values: 8000.0 m³ (mean), 400.0 m³ (std)
Relative error: 0.0% (mean), 3.2% (std)
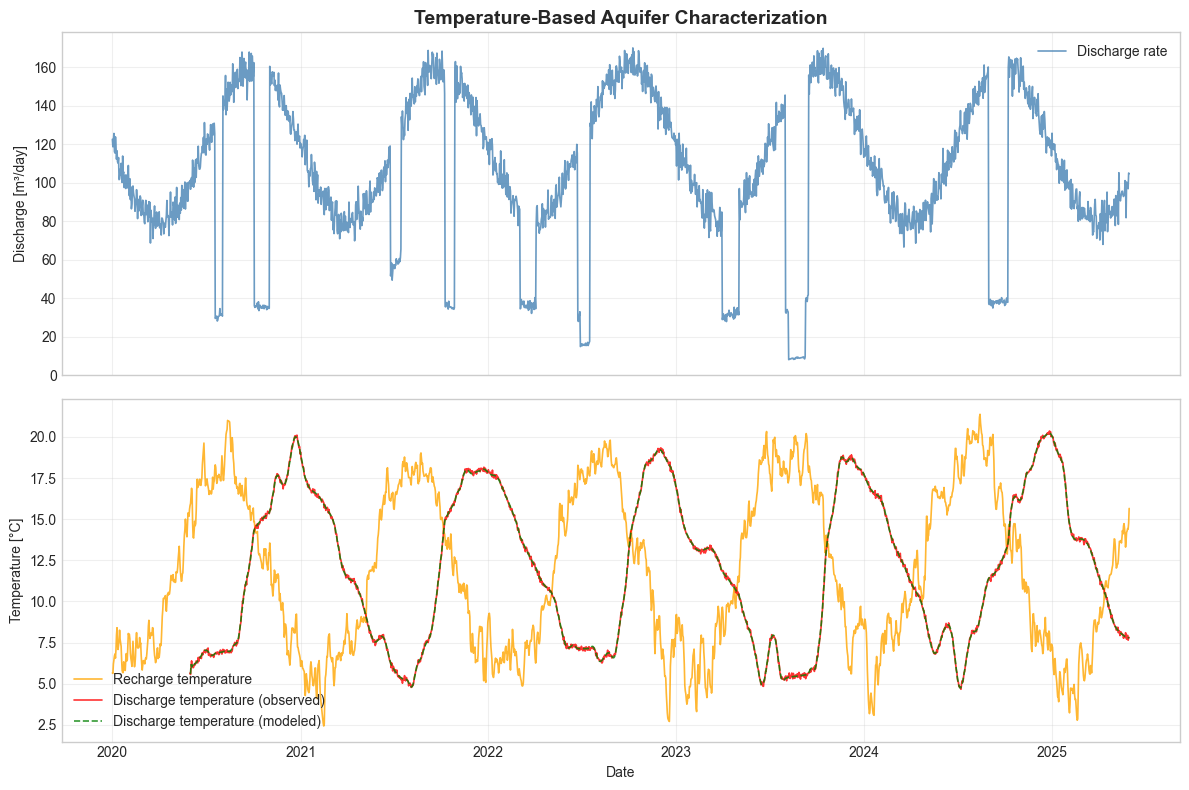
3. Model Validation and Visualization#
We compare observed and modeled temperature breakthrough curves to assess model performance.
[7]:
fig, (ax1, ax2) = plt.subplots(figsize=(12, 8), nrows=2, ncols=1, sharex=True)
# Flow rate subplot - convert to step format
xstep_flow, ystep_flow = step_plot_coords(tedges, df.flow)
ax1.set_title("Temperature-Based Aquifer Characterization", fontsize=14, fontweight="bold")
ax1.plot(xstep_flow, ystep_flow, label="Discharge rate", color="steelblue", alpha=0.8, linewidth=1.2)
ax1.set_ylabel("Discharge [m³/day]")
ax1.legend()
ax1.grid(True, alpha=0.3)
# Temperature subplot - convert all series to step format
xstep_temp, ystep_infiltration = step_plot_coords(tedges, df.cin)
_, ystep_extraction = step_plot_coords(tedges, df.cout)
_, ystep_modeled = step_plot_coords(tedges, df.cout_modeled)
ax2.plot(xstep_temp, ystep_infiltration, label="Recharge temperature", color="orange", alpha=0.8, linewidth=1.2)
ax2.plot(xstep_temp, ystep_extraction, label="Discharge temperature (observed)", color="red", alpha=0.8, linewidth=1.2)
ax2.plot(
xstep_temp,
ystep_modeled,
label="Discharge temperature (modeled)",
color="green",
alpha=0.8,
linewidth=1.2,
linestyle="--",
)
ax2.set_xlabel("Date")
ax2.set_ylabel("Temperature [°C]")
ax2.legend()
ax2.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
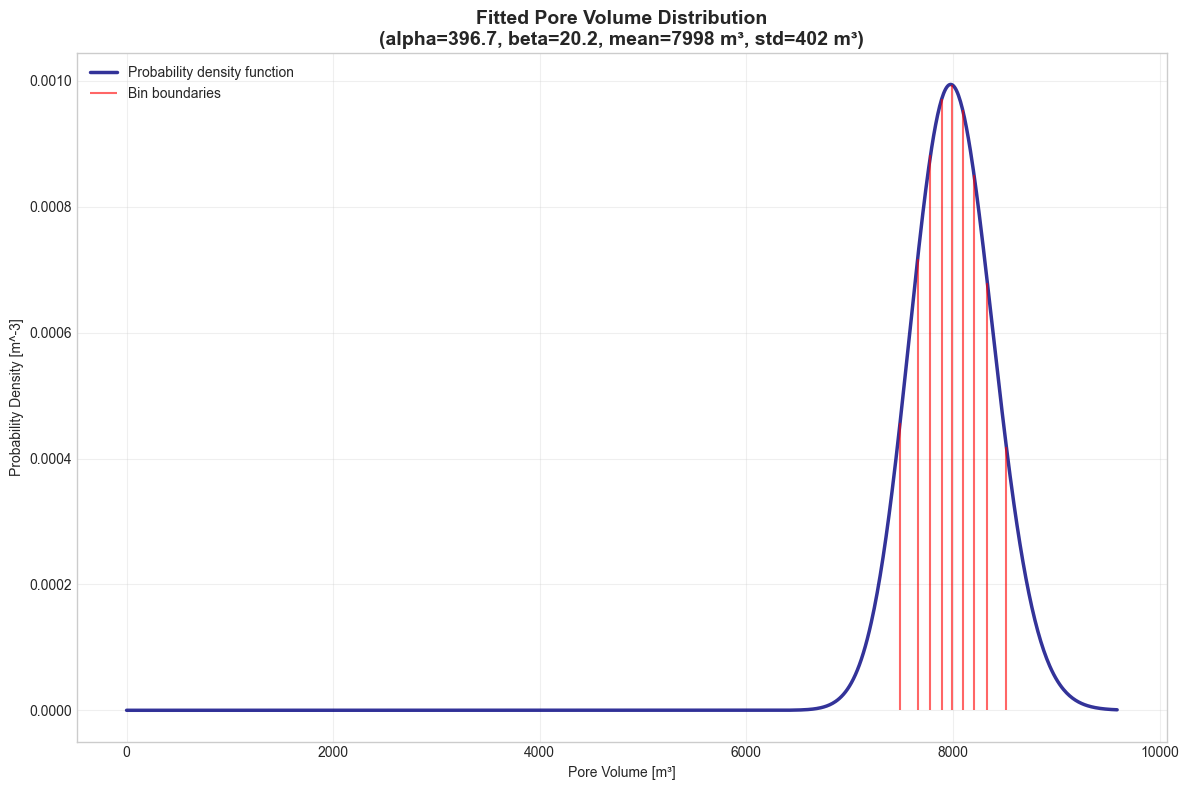
4. Pore Volume Distribution Analysis#
We visualize the fitted gamma distribution representing spatial heterogeneity in pore volume. Each bin represents a different flow path through the aquifer.
[8]:
# Discretize gamma distribution into flow path bins
n_bins = 10 # Reduced for visualization clarity
alpha, beta = gamma_utils.mean_std_loc_to_alpha_beta(mean=mean, std=std)
gbins = gamma_utils.bins(alpha=alpha, beta=beta, n_bins=n_bins)
print(f"Gamma Distribution Parameters: alpha={alpha:.1f}, beta={beta:.1f}")
print(f"Discretized into {n_bins} equiprobable bins:")
print("-" * 80)
print(f"{'Bin':3s} {'Lower [m³]':10s} {'Upper [m³]':10s} {'E[V|bin]':10s} {'P(bin)':10s}")
print("-" * 80)
for i in range(n_bins):
upper = f"{gbins['upper_bound'][i]:.1f}" if not np.isinf(gbins["upper_bound"][i]) else "∞"
lower = f"{gbins['lower_bound'][i]:.1f}"
expected = f"{gbins['expected_values'][i]:.1f}"
prob = f"{gbins['probability_mass'][i]:.3f}"
print(f"{i:3d} {lower:10s} {upper:10s} {expected:10s} {prob:10s}")
Gamma Distribution Parameters: alpha=426.9, beta=18.7
Discretized into 10 equiprobable bins:
--------------------------------------------------------------------------------
Bin Lower [m³] Upper [m³] E[V|bin] P(bin)
--------------------------------------------------------------------------------
0 0.0 7510.3 7337.0 0.100
1 7510.3 7674.9 7598.7 0.100
2 7674.9 7795.0 7737.0 0.100
3 7795.0 7898.6 7847.6 0.100
4 7898.6 7996.3 7947.6 0.100
5 7996.3 8094.8 8045.2 0.100
6 8094.8 8201.0 8146.9 0.100
7 8201.0 8326.5 8261.4 0.100
8 8326.5 8502.7 8407.6 0.100
9 8502.7 ∞ 8696.2 0.100
[9]:
# Plot the gamma distribution and bins
x = np.linspace(0, 1.1 * gbins["expected_values"][-1], 1000)
y = gamma_dist.pdf(x, alpha, scale=beta)
fig, ax = plt.subplots(figsize=(12, 8))
ax.set_title(
f"Fitted Pore Volume Distribution\n(alpha={alpha:.1f}, beta={beta:.1f}, mean={mean:.0f} m³, std={std:.0f} m³)",
fontsize=14,
fontweight="bold",
)
ax.plot(x, y, label="Probability density function", color="navy", alpha=0.8, linewidth=2.5)
pdf_at_lower_bound = gamma_dist.pdf(gbins["lower_bound"], alpha, scale=beta)
ax.vlines(
gbins["lower_bound"],
0,
pdf_at_lower_bound,
color="red",
alpha=0.6,
linewidth=1.5,
label="Bin boundaries",
)
ax.set_xlabel("Pore Volume [m³]")
ax.set_ylabel("Probability Density [m^-3]")
ax.legend()
ax.grid(True, alpha=0.3)
plt.tight_layout()
Results & Discussion#
The inverse modeling successfully recovered the aquifer pore volume distribution parameters. The fitted gamma distribution captures the heterogeneity in flow paths through the aquifer.
Practical Applications#
Contaminant Transport: Use fitted parameters to predict pollutant breakthrough
Well Field Design: Optimize extraction rates based on residence time requirements
Vulnerability Assessment: Identify fast flow paths that may compromise water quality
Key Takeaways#
Temperature as Natural Tracer: Temperature provides valuable information about aquifer properties without artificial injection
Inverse Modeling: Optimization techniques can extract quantitative aquifer parameters from field observations
Gamma Distribution: Effective model for representing aquifer heterogeneity in engineering applications
Thermal Retardation: Must account for heat exchange when using temperature data
Spin-up Period: Use :func:
~gwtransport.residence_time.fraction_explainedto determine when the model output is fully informed by the input signal