Note
This notebook is located in the ./examples directory of the gwtransport repository.
Pathogen Removal in Bank Filtration Systems#
Learning Objectives#
Understand log-removal concepts for pathogen treatment assessment
Calculate pathogen removal efficiency in groundwater treatment systems
Learn how heterogeneous systems affect overall performance
Apply residence time analysis to water treatment design
Analyze seasonal variations in treatment performance
Overview#
This notebook demonstrates how to calculate pathogen removal efficiency in bank filtration systems using log-removal analysis. River water infiltrates through riverbank sediments, where pathogens are removed through biological decay (inactivation) and physical straining (attachment).
Key Concepts#
Log-removal: Logarithmic scale for pathogen reduction (1 log = 90%, 2 log = 99%, 3 log = 99.9%)
Two-component model: Total removal = inactivation (time-dependent) + attachment (geometry-dependent)
Heterogeneous systems: Overall performance weighted toward worst-performing flow paths
Background Reading#
Residence Time — Contact time determines removal efficiency
Pore Volume Distribution — Flow path heterogeneity affects overall performance
Theoretical Background#
Two-Component Pathogen Removal Model#
Total pathogen removal during soil passage consists of two mechanisms (Schijven and Hassanizadeh, 2000):
Inactivation (time-dependent): First-order biological decay: \(\text{LR}_\text{decay} = \mu \cdot t_{residence}\), where \(\mu\) is the log10 decay rate [log10/day]. Temperature-dependent: \(\ln(\mu_l) = a \cdot T + b\).
Attachment (geometry-dependent): Physical removal by adsorption and straining. Depends on aquifer geometry, travel distance, and soil properties. At Castricum, Schijven et al. (1999) found attachment contributed ~97% of total MS2 removal.
The gwtransport.logremoval module models the time-dependent decay component. Users add geometry-dependent attachment based on site-specific knowledge.
Decay Rate Conversion#
The natural-log decay rate \(\lambda\) [1/day] relates to the log10 rate \(\mu\) by: \(\mu = \lambda / \ln(10)\). Use decay_rate_to_log10_decay_rate() to convert published rates.
See residence time for background on how contact time determines removal efficiency.
[1]:
import matplotlib.pyplot as plt
import numpy as np
from gwtransport import gamma as gamma_utils
from gwtransport.examples import generate_temperature_example_data
from gwtransport.logremoval import (
decay_rate_to_log10_decay_rate,
gamma_find_flow_for_target_mean,
gamma_mean,
parallel_mean,
residence_time_to_log_removal,
)
from gwtransport.residence_time import fraction_explained, residence_time
from gwtransport.utils import step_plot_coords
# Set random seed for reproducibility
np.random.seed(42)
plt.style.use("seaborn-v0_8-whitegrid")
print("Libraries imported successfully")
Libraries imported successfully
1. Understanding Pathogen Inactivation Rates#
Pathogen inactivation rates vary significantly between virus types and temperatures. Schijven and Hassanizadeh (2000) compiled inactivation rates from multiple studies and found that the temperature dependence follows: ln(mu_l) = a*T + b, where T is the groundwater temperature in degrees Celsius.
The table below shows inactivation rates at 10 degrees C for common model viruses:
[2]:
print("=== Pathogen Inactivation Rates ===")
print("From Schijven and Hassanizadeh (2000), Table 7")
print("Temperature regression: ln(mu_l) = a*T + b\n")
# Inactivation rate regression parameters from Schijven & Hassanizadeh (2000) Table 7
# mu_l is the natural-log inactivation rate [day^-1]
# ln(mu_l) = a * T + b, where T is temperature in degrees C
pathogen_params = {
"MS2": {"a": 0.12, "b": -3.5, "R2": 0.85},
"PRD1": {"a": 0.09, "b": -4.0, "R2": 0.71},
"Poliovirus 1": {"a": 0.033, "b": -2.4, "R2": 0.07},
"Echovirus 1": {"a": 0.12, "b": -3.0, "R2": 0.71},
"HAV": {"a": -0.024, "b": -1.4, "R2": 0.08},
}
def inactivation_rate_log10(temperature, a, b):
"""Compute log10 inactivation rate [log10/day] at temperature [deg C]."""
mu_l = np.exp(a * temperature + b) # natural-log rate [1/day]
return mu_l / np.log(10) # convert to log10 rate [log10/day]
# Evaluate at T = 10 deg C (typical Dutch groundwater)
T_ref = 10.0
print(f"Inactivation rates at T = {T_ref:.0f} deg C:")
print(f"{'Virus':<16} {'mu_l [1/day]':>12} {'mu [log10/day]':>15} {'Temp. sensitivity':>18}")
print("-" * 65)
log10_decay_rates = {}
for virus, params in pathogen_params.items():
mu_l = np.exp(params["a"] * T_ref + params["b"])
mu_log10 = mu_l / np.log(10)
log10_decay_rates[virus] = mu_log10
r2_str = f"R2={params['R2']:.0%}" if params["R2"] > 0.5 else f"R2={params['R2']:.0%} (weak)"
print(f" {virus:<14} {mu_l:>12.4f} {mu_log10:>15.4f} {r2_str:>18}")
print("\nNote: These are INACTIVATION rates only (time-dependent decay).")
print("Total pathogen removal also includes attachment (geometry-dependent),")
print("which typically dominates and is not modeled by gwtransport.")
=== Pathogen Inactivation Rates ===
From Schijven and Hassanizadeh (2000), Table 7
Temperature regression: ln(mu_l) = a*T + b
Inactivation rates at T = 10 deg C:
Virus mu_l [1/day] mu [log10/day] Temp. sensitivity
-----------------------------------------------------------------
MS2 0.1003 0.0435 R2=85%
PRD1 0.0450 0.0196 R2=71%
Poliovirus 1 0.1262 0.0548 R2=7% (weak)
Echovirus 1 0.1653 0.0718 R2=71%
HAV 0.1940 0.0842 R2=8% (weak)
Note: These are INACTIVATION rates only (time-dependent decay).
Total pathogen removal also includes attachment (geometry-dependent),
which typically dominates and is not modeled by gwtransport.
[3]:
print("=== Converting Published Decay Rates ===")
print("Example: MS2 inactivation rate from Schijven et al. (1999) Castricum field study\n")
# MS2 free virus inactivation rate at 5 deg C from Castricum field study
# Published as natural-log rate: mu_f = 0.03 day^-1
published_decay_rate = 0.03 # 1/day (natural-log inactivation rate for MS2 at 5 deg C)
# Convert to log10 decay rate
converted_rate = decay_rate_to_log10_decay_rate(published_decay_rate)
print(f"Published natural-log inactivation rate (lambda): {published_decay_rate} 1/day")
print(f"Converted log10 decay rate (mu): {converted_rate:.4f} log10/day")
print(f"\nVerification: mu * ln(10) = {converted_rate * np.log(10):.4f} (should equal lambda)")
# Compare with regression estimate
t_ref = 5.0
regression_rate = inactivation_rate_log10(t_ref, **{k: v for k, v in pathogen_params["MS2"].items() if k != "R2"})
print(f"\nRegression estimate for MS2 at {t_ref:.0f} deg C: {regression_rate:.4f} log10/day")
print(f"Field measurement at {t_ref:.0f} deg C: {converted_rate:.4f} log10/day")
# Show what this means for a typical residence time
mean_residence_time = 76.0 # days (example from article)
print(f"\nFor a mean residence time of {mean_residence_time:.0f} days:")
print(
f" LR_decay (inactivation only) = {converted_rate:.4f} * {mean_residence_time:.0f} = {converted_rate * mean_residence_time:.2f} log10"
)
print(" This is just the time-dependent component.")
print(" The user must add site-specific attachment removal separately.")
=== Converting Published Decay Rates ===
Example: MS2 inactivation rate from Schijven et al. (1999) Castricum field study
Published natural-log inactivation rate (lambda): 0.03 1/day
Converted log10 decay rate (mu): 0.0130 log10/day
Verification: mu * ln(10) = 0.0300 (should equal lambda)
Regression estimate for MS2 at 5 deg C: 0.0239 log10/day
Field measurement at 5 deg C: 0.0130 log10/day
For a mean residence time of 76 days:
LR_decay (inactivation only) = 0.0130 * 76 = 0.99 log10
This is just the time-dependent component.
The user must add site-specific attachment removal separately.
2. Heterogeneous System Performance#
Bank filtration systems have multiple flow paths with different residence times. The overall log-removal is weighted toward lower values (shorter residence times).
[4]:
print("=== Heterogeneous System Analysis ===")
print("Multiple flow paths with different residence times\n")
# Three flow paths with different log-removal efficiencies
unit_removals = np.array([0.5, 1.0, 1.5]) # log10 values for each path
# Correct method: parallel_mean() accounts for flow-weighted averaging
combined_removal = parallel_mean(log_removals=unit_removals)
print("Flow Path Performance:")
for i, removal in enumerate(unit_removals):
efficiency = (1 - 10 ** (-removal)) * 100
print(f" Path {i + 1}: {removal:.1f} log10 -> {efficiency:.1f}% removal")
print("\nOverall System Performance:")
combined_efficiency = (1 - 10 ** (-combined_removal)) * 100
print(f" Combined log-removal: {combined_removal:.2f} log10 -> {combined_efficiency:.1f}% removal")
print("\nNote: Overall performance is weighted toward the worst-performing paths")
print("(shortest residence times), ensuring conservative design.")
=== Heterogeneous System Analysis ===
Multiple flow paths with different residence times
Flow Path Performance:
Path 1: 0.5 log10 -> 68.4% removal
Path 2: 1.0 log10 -> 90.0% removal
Path 3: 1.5 log10 -> 96.8% removal
Overall System Performance:
Combined log-removal: 0.83 log10 -> 85.1% removal
Note: Overall performance is weighted toward the worst-performing paths
(shortest residence times), ensuring conservative design.
3. Design Application - Achieving Target Removal Efficiency#
Water treatment facilities aim to achieve specific pathogen removal targets. We demonstrate how to design systems that achieve desired removal efficiency.
[5]:
print("=== Design Application ===")
print("Design challenge: Achieve target inactivation for safe drinking water\n")
# Define aquifer characteristics (small riverbank aquifer)
mean_pore_volume = 1000.0 # m3 (total water-filled space)
std_pore_volume = 300.0 # m3 (variability in pore volume)
flow_rate = 50.0 # m3/day (water extraction rate)
# Convert aquifer properties to gamma distribution parameters
alpha, beta = gamma_utils.mean_std_loc_to_alpha_beta(mean=mean_pore_volume, std=std_pore_volume)
# Use MS2 at 10 deg C as the design pathogen (worst-case model virus)
log10_decay_rate = log10_decay_rates["MS2"] # ~0.043 log10/day
# Target inactivation level (decay component only)
target_decay = 3.0 # log10 inactivation target
print(f"Pathogen: MS2 at 10 deg C (mu = {log10_decay_rate:.4f} log10/day)")
print(f"Target inactivation (decay only): {target_decay} log10")
# Calculate effective decay at current flow
rt_alpha = alpha
rt_beta = beta / flow_rate
mean_log_removal = gamma_mean(rt_alpha=rt_alpha, rt_beta=rt_beta, log10_decay_rate=log10_decay_rate)
mean_residence_time = mean_pore_volume / flow_rate
print(f"\nAt flow rate {flow_rate} m3/day (mean RT = {mean_residence_time:.0f} days):")
print(f" Effective inactivation: {mean_log_removal:.2f} log10")
# Find the maximum flow rate that still achieves the inactivation target
required_flow = gamma_find_flow_for_target_mean(
target_mean=target_decay,
apv_alpha=alpha,
apv_beta=beta,
log10_decay_rate=log10_decay_rate,
)
required_residence_time = mean_pore_volume / required_flow
print(f"\nMaximum flow for {target_decay} log10 inactivation target:")
print(f" Maximum flow rate: {required_flow:.1f} m3/day")
print(f" Required mean residence time: {required_residence_time:.1f} days")
# Compare all pathogens
print(f"\nMaximum flow rates for {target_decay} log10 inactivation target (all pathogens):")
for virus, mu in log10_decay_rates.items():
max_flow = gamma_find_flow_for_target_mean(
target_mean=target_decay,
apv_alpha=alpha,
apv_beta=beta,
log10_decay_rate=mu,
)
print(f" {virus:<16}: {max_flow:>6.1f} m3/day (mu = {mu:.4f} log10/day)")
=== Design Application ===
Design challenge: Achieve target inactivation for safe drinking water
Pathogen: MS2 at 10 deg C (mu = 0.0435 log10/day)
Target inactivation (decay only): 3.0 log10
At flow rate 50.0 m3/day (mean RT = 20 days):
Effective inactivation: 0.80 log10
Maximum flow for 3.0 log10 inactivation target:
Maximum flow rate: 10.5 m3/day
Required mean residence time: 95.5 days
Maximum flow rates for 3.0 log10 inactivation target (all pathogens):
MS2 : 10.5 m3/day (mu = 0.0435 log10/day)
PRD1 : 4.7 m3/day (mu = 0.0196 log10/day)
Poliovirus 1 : 13.2 m3/day (mu = 0.0548 log10/day)
Echovirus 1 : 17.3 m3/day (mu = 0.0718 log10/day)
HAV : 20.3 m3/day (mu = 0.0842 log10/day)
4. Real-World Scenario - Seasonal Variations#
In reality, river flows change seasonally, affecting bank filtration performance. We simulate a multi-year system to see how log-removal varies with changing conditions.
[6]:
print("=== Seasonal Flow Data Generation ===")
# Generate realistic flow data with seasonal patterns
df, tedges = generate_temperature_example_data(
date_start="2020-01-01",
date_end="2025-05-31",
flow_mean=120.0, # Base flow rate [m³/day]
flow_amplitude=40.0, # Seasonal flow variation [m³/day]
flow_noise=5.0, # Random daily fluctuations [m³/day]
cin_method="soil_temperature", # Use real soil temperature data
aquifer_pore_volume_gamma_mean=8000.0, # True mean pore volume [m³]
aquifer_pore_volume_gamma_std=400.0, # True standard deviation [m³]
)
# Set up aquifer characteristics for this larger system
bins = gamma_utils.bins(
mean=df.attrs["aquifer_pore_volume_gamma_mean"], std=df.attrs["aquifer_pore_volume_gamma_std"], n_bins=1000
) # High resolution
print(f"Dataset: {len(df)} days from {df.index[0].date()} to {df.index[-1].date()}")
print(f"Flow range: {df['flow'].min():.1f} - {df['flow'].max():.1f} m³/day")
print(
f"Aquifer: {df.attrs['aquifer_pore_volume_gamma_mean']:.0f} ± {df.attrs['aquifer_pore_volume_gamma_std']:.0f} m³ pore volume"
)
=== Seasonal Flow Data Generation ===
Dataset: 1978 days from 2020-01-01 to 2025-05-31
Flow range: 6.4 - 170.8 m³/day
Aquifer: 8000 ± 400 m³ pore volume
[7]:
print("\nComputing inactivation for multiple pathogens over time...")
# Calculate residence time distribution for water flow
rt_infiltration_to_extraction_water = residence_time(
flow=df.flow,
flow_tedges=tedges,
aquifer_pore_volumes=bins["expected_values"],
retardation_factor=1.0, # Water (conservative tracer)
direction="infiltration_to_extraction",
)
# Compute fraction explained to assess spin-up
frac = fraction_explained(rt=rt_infiltration_to_extraction_water)
fully_explained_mask = frac >= 1.0
if fully_explained_mask.any():
print(f"Spin-up period ends: {df.index[fully_explained_mask][0].date()}")
print(" Log-removal values during spin-up (fraction explained < 1.0) are unreliable")
# Compute log-removal for each pathogen
for virus, mu in log10_decay_rates.items():
col = f"lr_decay_{virus}"
log_removal_array = residence_time_to_log_removal(
residence_times=rt_infiltration_to_extraction_water,
log10_decay_rate=mu,
)
df[col] = parallel_mean(log_removals=log_removal_array, axis=0)
print("Time series calculation completed for all pathogens")
print("\nInactivation ranges (decay component only, fully explained period):")
for virus in log10_decay_rates:
col = f"lr_decay_{virus}"
valid = df.loc[fully_explained_mask, col]
print(f" {virus:<16}: {valid.min():.2f} - {valid.max():.2f} log10")
Computing inactivation for multiple pathogens over time...
Spin-up period ends: 2020-01-01
Log-removal values during spin-up (fraction explained < 1.0) are unreliable
Time series calculation completed for all pathogens
Inactivation ranges (decay component only, fully explained period):
MS2 : 2.17 - 6.18 log10
PRD1 : 0.98 - 2.79 log10
Poliovirus 1 : 2.72 - 7.76 log10
Echovirus 1 : 3.56 - 10.12 log10
HAV : 4.17 - 11.84 log10
5. Performance Visualization#
[8]:
# Create time series plot showing inactivation for multiple pathogens
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(14, 10), sharex=True)
# Plot 1: Flow rate over time
ax1.plot(*step_plot_coords(tedges, df.flow), color="steelblue", linewidth=1.2, alpha=0.8)
ax1.set_ylabel("Flow Rate (m3/day)")
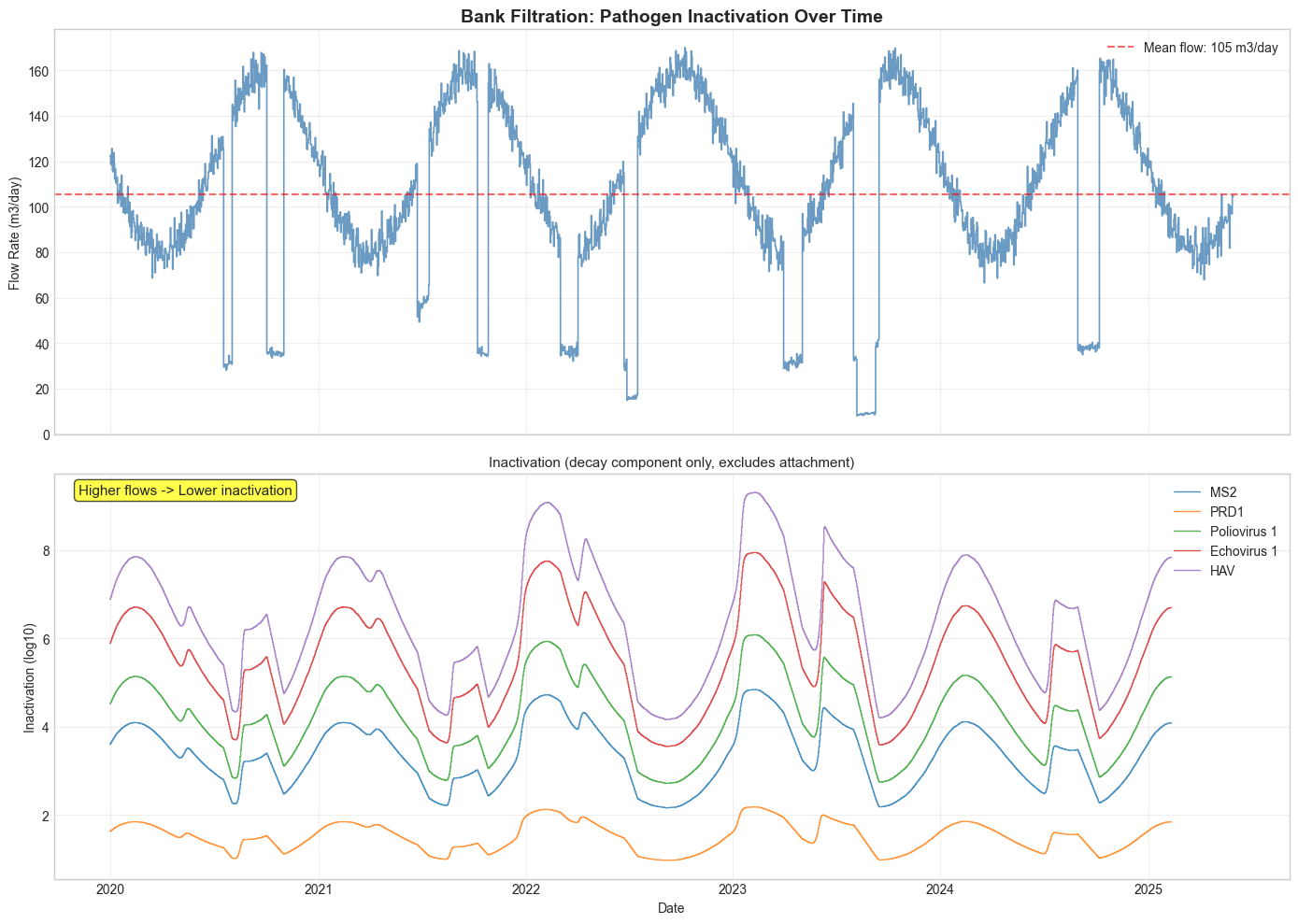
ax1.set_title("Bank Filtration: Pathogen Inactivation Over Time", fontsize=14, fontweight="bold")
ax1.grid(True, alpha=0.3)
ax1.axhline(
y=df.flow.mean(),
color="red",
linestyle="--",
alpha=0.6,
label=f"Mean flow: {df.flow.mean():.0f} m3/day",
)
ax1.legend()
# Plot 2: Inactivation for each pathogen
colors = {"MS2": "C0", "PRD1": "C1", "Poliovirus 1": "C2", "Echovirus 1": "C3", "HAV": "C4"}
for virus in log10_decay_rates:
col = f"lr_decay_{virus}"
ax2.plot(*step_plot_coords(tedges, df[col]), color=colors[virus], linewidth=1.0, alpha=0.8, label=virus)
ax2.set_ylabel("Inactivation (log10)")
ax2.set_xlabel("Date")
ax2.grid(True, alpha=0.3)
ax2.legend(loc="upper right")
ax2.set_title("Inactivation (decay component only, excludes attachment)", fontsize=11)
# Add fraction explained on twin axis
ax2b = ax2.twinx()
ax2b.plot(
*step_plot_coords(tedges, frac * 100),
color="gray",
linewidth=1.5,
linestyle=":",
alpha=0.8,
label="Fraction explained",
)
ax2b.set_ylabel("Fraction Explained [%]")
ax2b.set_ylim(0, 105)
ax2b.legend(loc="lower right", fontsize=9)
ax2.text(
0.02,
0.95,
"Higher flows -> Lower inactivation",
transform=ax2.transAxes,
fontsize=11,
bbox={"boxstyle": "round,pad=0.3", "facecolor": "yellow", "alpha": 0.7},
)
plt.tight_layout()
plt.show()
6. Performance Summary and Analysis#
[9]:
# Summary statistics for all pathogens (fully explained period only)
print("Performance Summary (inactivation component only, fully explained period):")
print("=" * 65)
for virus, mu in log10_decay_rates.items():
col = f"lr_decay_{virus}"
valid = df.loc[fully_explained_mask, col]
min_lr = valid.min()
max_lr = valid.max()
mean_lr = valid.mean()
std_lr = valid.std()
print(f"\n{virus} (mu = {mu:.4f} log10/day at 10 deg C):")
print(f" Range: {min_lr:.2f} - {max_lr:.2f} log10")
print(f" Mean: {mean_lr:.2f} +/- {std_lr:.2f} log10")
print("\n" + "=" * 65)
print("\nRemember: Total pathogen removal = inactivation + attachment.")
print("The attachment component (geometry-dependent) must be added separately")
print("based on site-specific data. At the Castricum site (Schijven et al., 1999),")
print("attachment contributed ~97% of total MS2 removal.")
Performance Summary (inactivation component only, fully explained period):
=================================================================
MS2 (mu = 0.0435 log10/day at 10 deg C):
Range: 2.17 - 6.18 log10
Mean: 3.41 +/- 0.84 log10
PRD1 (mu = 0.0196 log10/day at 10 deg C):
Range: 0.98 - 2.79 log10
Mean: 1.54 +/- 0.38 log10
Poliovirus 1 (mu = 0.0548 log10/day at 10 deg C):
Range: 2.72 - 7.76 log10
Mean: 4.29 +/- 1.06 log10
Echovirus 1 (mu = 0.0718 log10/day at 10 deg C):
Range: 3.56 - 10.12 log10
Mean: 5.59 +/- 1.38 log10
HAV (mu = 0.0842 log10/day at 10 deg C):
Range: 4.17 - 11.84 log10
Mean: 6.54 +/- 1.61 log10
=================================================================
Remember: Total pathogen removal = inactivation + attachment.
The attachment component (geometry-dependent) must be added separately
based on site-specific data. At the Castricum site (Schijven et al., 1999),
attachment contributed ~97% of total MS2 removal.
Results & Discussion#
Seasonal Performance#
High flow periods reduce residence times, leading to lower log-removal. Systems must be designed for worst-case (high flow) conditions.
Heterogeneous System Behavior#
Overall performance is weighted toward shortest residence times (worst-performing flow paths), providing natural safety margins in design.
Engineering Design#
Trade-off between water production and treatment efficiency
Design for worst-case: high flow (short RT) + low temperature (low decay rate)
Monitor performance during high-flow periods
Key Takeaways#
Two-Component Model: Total removal = inactivation (time-dependent, modeled by gwtransport) + attachment (geometry-dependent, user-specified)
Residence Time Dependency: Longer underground residence = better pathogen removal
Flow Rate Trade-off: Higher pumping rates reduce treatment efficiency
Heterogeneous Systems: Overall performance weighted toward worst-performing flow paths
Design for Worst Case: High flow + low temperature = minimum removal
Engineering Design Summary#
Inactivation rates: Use
decay_rate_to_log10_decay_rate()to convert published natural-log rates. Key reference: Schijven and Hassanizadeh (2000) Table 7. MS2 is the standard worst-case model virus.Maximum safe flow: Use
gamma_find_flow_for_target_mean()to find the maximum flow rate achieving a target inactivation level.Temperature dependence: Design for winter conditions (lowest temperatures, lowest inactivation rates). Regression: \(\ln(\mu_l) = a \cdot T + b\).